How Φ80 Infiltrates Research Labs
While some bacteriophages play vital roles in laboratory research, others are bent on sabotage.
Living organisms are surprisingly creative when it comes to finding their way into hostile environments. Microbes, after all, thrive everywhere from hyperarid deserts and hydrothermal vents to the stratosphere.
Of all hostile environments, one might think the most unbreachable to be the laboratory, where populations of bacteria are stored inside tubes sealed with tight screw-caps and stacked in freezers at -80°C. Of course, normal evolutionary processes don’t magically stop at the doors of the lab: organisms still mutate, fragile microbial species can still be out-competed by faster-growing contaminants, and parasites can still spread. However, vigilant scientists devote considerable effort to making sure containers hold what is written on their labels and, when in doubt, can sequence any part of a sample’s DNA, discarding or incinerating undesired specimens.
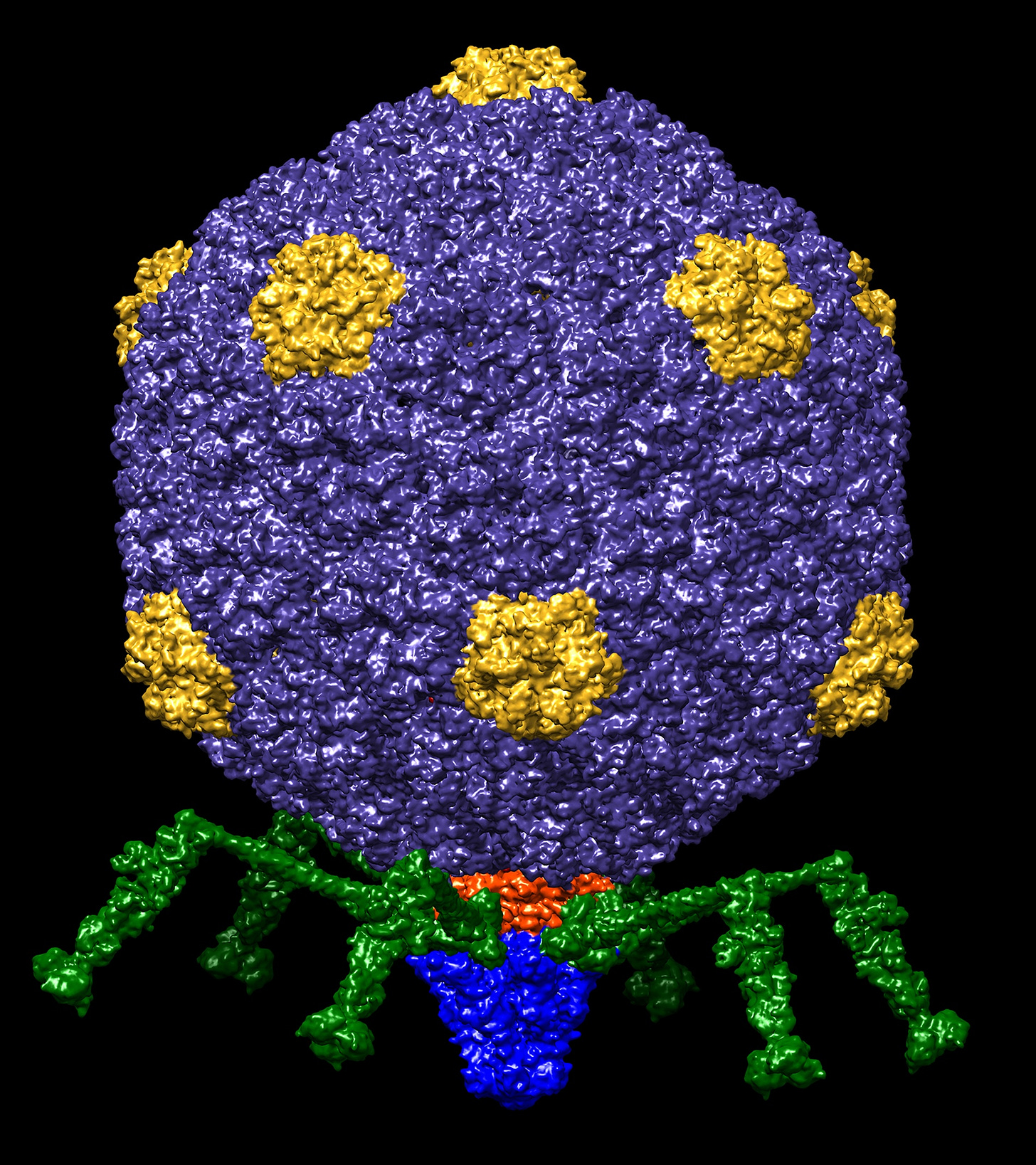
Still, interlopers persist. One organism that has been particularly successful at infiltrating microbiology research facilities is called Φ80. It is a bacteriophage, or virus that infects bacteria. Slowly but surely, Φ80 has been spreading. Researchers carry out experiments, make reproducible observations, and publish — usually without ever realizing that their samples have quietly been contaminated with this phage.
On occasion, hidden contaminations of Φ80 have led to paper retractions, with researchers discovering not only that their results are artifactual, but that they personally amplified the invasion and transmitted it elsewhere. In a 2017 paper, for example, scientists confessed that “beyond the confusion created by our erroneous interpretation of results obtained with Φ80-infected strains, we are worried that we have sent these strains to many research groups.”
To understand how such unwitting contamination can occur despite researchers’ best efforts, and to work towards better containment, it helps to first understand how bacteria are typically cultivated.
Life in a Tube
The most commonly studied bacterium, Escherichia coli (E. coli), has spent most of its recent evolution inside the human gut. Acclimated to that environment, E. coli grows best at 37°C — dividing about every 20 minutes in good conditions. If a researcher inoculates just a few E. coli cells in a rich liquid medium and puts the tube in a shaking incubator, it only takes about a dozen hours to form a population of several billion cells. As these billions of cells continue to divide every 20 minutes, their consumption of nutrients also grows exponentially, eventually exhausting all resources, at which point they enter a kind of dormant state.
E. coli cells are almost completely transparent, but their refractive index differs slightly from the medium in which they are grown. Each cell is only about one micrometer wide, but when millions accumulate in the same test tube, visible light gets scattered, and the liquid becomes opaque.
Compared to animal guts, with their folds, crypts, and periodic floods of bile, a test tube is, from the point of view of a bacterium, a featureless space. Most importantly, it is also the only place on Earth where bacteria don’t have to deal with phages. While we don’t pay much attention to phages, they are everywhere, and Earth would be quite different without them. Every day, phages kill 15 to 40 percent of all bacteria in the ocean, keeping microbial ecosystems from spiraling out of balance.
A thriving culture of bacteria, peacefully growing in a test tube, would surely be an appetizing ground for a bacteria-killing phage to replicate, if it could find its way inside. For this reason, microbiologists are extremely vigilant about phage contamination. Merely mentioning phages at lab meetings will draw nervous looks!
Phages get everywhere, are hard to destroy, and tear through experiments. I’ve seen entire labs shut down for days while whole HVAC systems are sterilized. If there is any detectable sign of unwanted phages, then, researchers stop what they’re doing, autoclave every object possible at the highest possible temperature, and wash the rest with the harshest chemicals available. While many bacteriophages are capable of infecting common laboratory strains, they can usually only spread so far.
The most straightforward strategy by which phages spread is by replicating as fast as they can, infecting as many cells as possible, and killing them in the process. Phages that employ this strategy are called lytic.
One example is T7, a lytic phage that infects E. coli. When T7 enters a bacterium, it uses the cell’s resources to generate copies of itself. This takes about 17 minutes, at which point the cell contains hundreds of phage particles. The phages then burst out of the cell in an event called lysis, searching for new cells to infect. The speed of this process depends on the population density of the bacteria. In ideal conditions, a phage population can expand by a factor of 10⁵–10⁶ every hour.
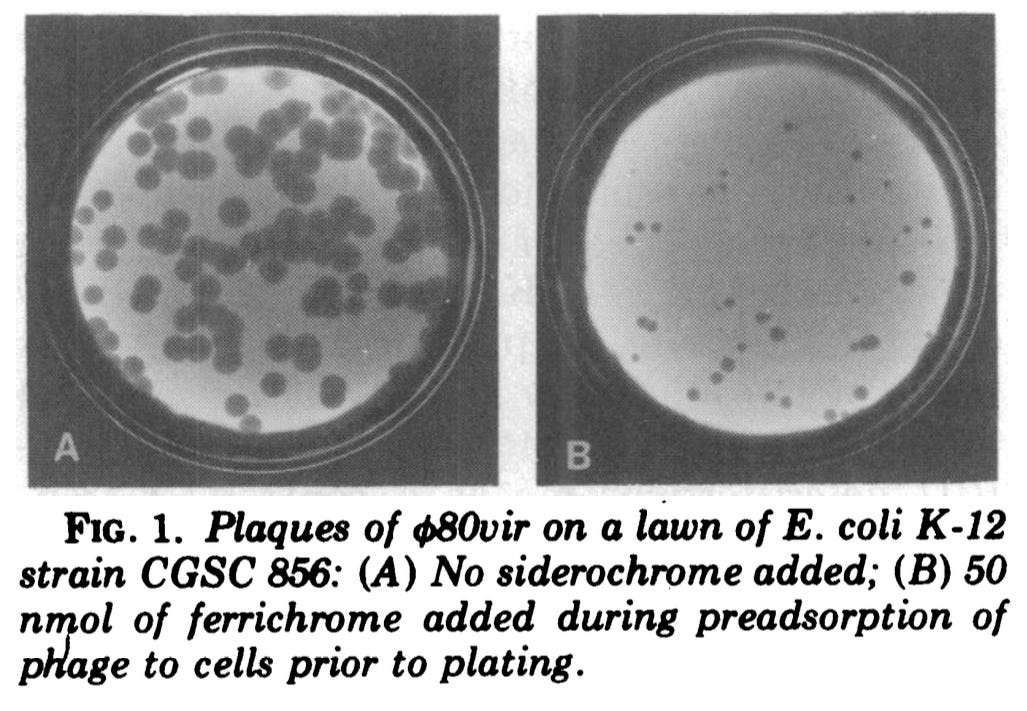
For bacteria in a test tube, phage infection means annihilation. Billions of cells can get entirely wiped out in two hours. As the bacteria burst and release their contents in the medium, the liquid becomes entirely transparent again.
While the lytic strategy employed by T7 is superb for proliferation, it doesn’t work well for secretly invading a laboratory. If an entire bacteriological culture suddenly collapses for no reason, even the most absent-minded researcher is apt to notice that their cells have been contaminated.
In fact, being too destructive is also a problem in the wild: if a phage eradicates its host too quickly, there might be no bacteria left to infect, meaning death for the phage as well. For that reason, many phages have adopted a more complex life cycle than that of T7, one known as lysogeny.
Take Phage λ, one of the best-studied lysogenic phages. Whenever it infects a new cell, it uses a genetic circuit to choose between two outcomes: either it goes through the usual lytic cycle, filling the cell with virions until it bursts, or it inserts its genome into the bacterium’s DNA. The integrated form is called a prophage.
The host bacterium then carries the phage genome even as it grows and divides. HIV behaves similarly in humans, but the advantage of doing so is much more obvious in bacteria: with a host that divides every 20 minutes, it can be worthwhile for phages to hitchhike for several generations of bacterial division, giving the host time to proliferate on its behalf.
It is even common for lysogenic phages to confer some benefit on their host, helping the infected bacteria outcompete their uninfected sisters and maximizing the prevalence of the prophage in the population.1 There is an interesting puzzle here: if there were an easy way to improve the fitness of the host bacteria, surely the bacteria would already have found it through normal natural selection. How, then, can a parasite grant an advantage that the host wouldn’t have evolved on its own?
One way is to provide the bacteria with genes helpful in the short-term at the expense of long-term sustainability. For example, phage-provided genes may offer costly solutions to temporary problems, like immunity to other pathogens or resistance to specific stresses in the environment.
When the host bacterium is itself a pathogen, phages can benefit from making bacteria more virulent to their host, often against the interests of the bacteria themselves. Some of the world’s most horrific diseases, like diphtheria, cholera, and botulism, are due to bacteria releasing deadly toxins into their host’s body. However, these toxin genes don’t come from the bacteria — they are often poisoned gifts from lysogenic phages.
So when do these dormant lysogenic phages elect to attack? The trigger is usually some indication that the cell is imperiled, whether by DNA damage, oxidative stress, or antibiotics. Some phages even force their hosts to secrete molecules into the environment so they can coordinate their decisions with phages in other infected cells. At the signal, like rats abandoning a sinking ship, the prophage leaves the chromosome, replicates profusely, and bursts into the environment.
In the context of the lab, this often resembles lytic infection, with most cells bursting and becoming transparent. But then, a small subpopulation of bacteria survives, having integrated the phage into their DNA. The prophage then expresses a protein that blocks further infection by the same phage, creating an immune population of cells that resumes growth as if nothing happened.
So, by using lysogeny, a phage can stay hidden in the chromosome of bacteria for extended periods of time. However, in a research lab environment, this tactic can only succeed if the phage manages to integrate itself into bacteria without the researchers noticing. And this is precisely why Φ80 is so cunning.
The Intruder
A key feature of Φ80 is that it replicates very slowly. While T7 goes through a full lifecycle in less than 20 minutes, Φ80 takes 90 minutes. This is a more dramatic difference than it seems: growth rate compounds with each generation, meaning that T7 can produce 1010 new particles in two hours, while Φ80 makes barely a thousand in the same interval.
Moreover, while the mechanism Φ80 uses to choose between lytic replication and chromosomal integration has not been fully deciphered, it appears to favor integration far more often than comparable lysogenic phages. As a result, a new infection by Φ80 will not cause the bacterial culture to suddenly collapse and become transparent. Instead, the phage will slowly establish itself inside the population in a way that is invisible to human overseers.
Additionally, it takes very little for Φ80 to switch back into lysis mode. Whereas many prophages use DNA damage as a signal that the bacterium is about to die and it’s time to look for a new host, Φ80 is so sensitive that even the slightest hiccup in DNA polymerization (something that happens routinely when a cell replicates) can make it go lytic. As cells carrying the Φ80 prophage grow, a tiny fraction will constantly pour new phage particles into the medium. As a result, billions of these phages will have accumulated in the tube over a cycle of growth, even though the culture continues to look perfectly healthy to the naked eye.
What’s worse is that this phage is mostly active at temperatures below 37°C. But, 37°C being the standard temperature for E. coli, pretty much every measurement is done at it. A researcher could put the culture on a slide in a microscope heated to 37°C, and everything would look fine. But as they took the culture through the corridor between the shaker and the microscope, the temperature would drop a little bit, and that’s precisely when a fraction of the culture would burst into a barely-noticeable, steady stream of Φ80 particles. In other words, the phage replicates precisely when researchers are not watching.
Even so, escaping detection is only half of its strategy. Φ80 also has an ingenious mechanism to break free from its tube and travel to new samples by hitchhiking on a classic molecular biology technique, thus turning human researchers into unwitting dispersers.
Scientists as Phage Dispersers
The technique in question is called P1-transduction. It is a widely used method for copying and pasting genes from one strain of bacteria to another. To understand how it works, we need to meet yet another phage: P1.2
The key to P1’s utility lies in a quirk of phage biology. During their reproductive cycles, phages use the bacteria’s machinery to produce a massive amount of empty capsids, which then house copies of the phage’s genome. This maintains a little army of fresh phage particles ready to go out and infect the next generation of cells.
Occasionally, though, there’s a mistake in the process: a piece of the host bacteria’s genome gets incorporated into a capsid instead of the phage’s genome. While this normally happens only rarely, P1 makes this mistake unusually often, about 0.1 percent of the time.3
As the phage particles are liberated in the medium and infect other bacteria, the random bit of DNA from the original, infected cell ends up injected into a different one.

This recipient cell notices that there’s a piece of linear, double-stranded DNA floating around in the cytoplasm. (E. coli’s chromosome is circular, so the presence of linear DNA is abnormal — it usually means that the chromosome is broken). In response, the bacterium expresses a suite of enzymes that stitch homologous pieces of DNA together based on sequence similarity, hoping to reconstruct a complete, circular chromosome. Thus, if P1 takes a piece of DNA from the first bacterium, then injects it into a second bacterium, the transplanted piece of DNA will often end up incorporated into the recipient’s chromosome, overwriting the native sequence. This process is called transduction.
Transduction gives scientists a clever way to copy and paste genes from one bacterial strain into another. First, the researcher inserts an antibiotic-resistance gene in the chromosome of one bacterium, called the donor strain, next to the gene they want to move.4 Then, they culture this donor bacterium in liquid broth, grow it up, and add a bit of phage P1. The P1 phages infect the donor cells, filling them up with invisible phage particles, including a few that took on a fragment of the donor strain’s DNA.
Next, these phage particles can be used to infect the recipient. At this point, random fragments from the donor DNA will be injected into the recipient and replace the native sequence by recombination. Finally, a researcher spreads the culture over a Petri dish containing the antibiotic from the first step, along with a chemical that inhibits further infection by P1. At this point, only the cells that have received the right region of the donor’s DNA — the bit containing an antibiotic resistance gene — can survive. And so, overnight, the few cells that received the desired edit will grow to form visible colonies.
When a two milliliter culture of bacteria gets infected by P1, this quickly generates tens of billions of phage particles. 0.1 percent of them will have incorporated a piece of the host DNA by mistake, which means our tube will contain around ten million such mistakes. Each of these pieces of DNA is about 90,000 base pairs long (like the phage’s genome). So, the total amount of donor DNA (mis)incorporated by the phage population will be almost a trillion nucleotides. For comparison, the full genome of E. coli is only 5 million base pairs. Therefore, the phages in our tubes collectively contain the entire genome of the donor bacterium: in fact, hundreds of thousands of copies of it. Add this to a recipient culture with several billion cells, and it’s almost guaranteed that one of them will receive the desired segment and incorporate it in the right place.
This is what makes P1 transduction a quintessential tool for microbiology. Although it relies on a series of unlikely coincidences, the sheer number of particles that fit in a droplet of liquid means it somehow works out in the end. And as long as a genetic element has an antibiotic-resistance marker next to it, it can be easily moved from strain to strain. The most famous application of this is the Keio collection, a holding of thousands of E. coli strains, each with a different gene replaced with an antibiotic resistance marker.
This ease of use made P1-transduction a hugely popular technique, used daily by thousands of biologists around the globe. Unfortunately, its popularity also makes it a perfect conduit for a stealthy parasite like Φ80 to travel from strain to strain. This is because, by a twist of fate, Φ80 has figured out a way to “tag along” with P1 phages.
Consider a routine P1-transduction experiment, aimed at moving a gene from donor to recipient. The donor strain, unknown to the researcher, harbors the Φ80 prophage. When P1 is added to the culture, the population collapses after a few hours; the expected sign of successful phage production. But the arrival of P1 is detected by the dormant Φ80 prophages, which now replicate alongside P1 in massive quantities. The oblivious researcher infects the recipient strain with this mixed phage solution, selects for the antibiotic resistance marker, and sequences the target locus. Everything appears to have worked as expected (the desired gene has been successfully transferred), but the new strain now carries Φ80.
This is, in fact, how Φ80 was discovered in the first place. In the late 2000s, researchers at the University of Illinois in Urbana-Champaign were trying to understand how bacteria repaired their DNA after damage from UV light. They suspected that LigB, a ligase that ties strands of DNA together, might play a role.
To test this hypothesis, the researchers did a classic knock-out experiment: they used P1-transduction to remove the ligB gene, then put cultures of the original bacteria and the modified ones side by side and exposed them to UV light. After exposure, the cultures of the strain lacking LigB mostly collapsed, while the natural strain didn’t. From that, they drew the logical conclusion that LigB must somehow repair UV damage. And yet, their assumptions were all wrong; the LigB ligase has nothing to do with UV damage.
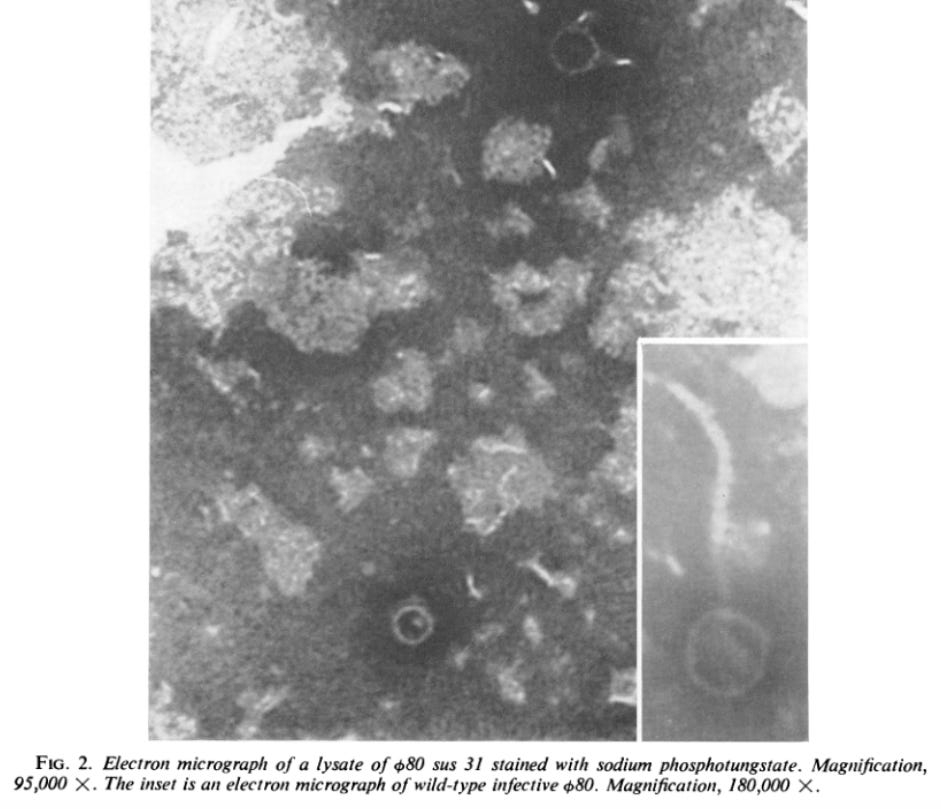
What really happened was that their donor strain was infected with Φ80, which took advantage of the P1-transduction to jump to the new strain. When this strain was exposed to UV light, the DNA damage induced Φ80’s lytic cycle, which looked exactly as if the cells were dying from the damage itself.
But why does P1 activate Φ80? Is it an ancestral relationship evolved over the course of millennia, or is it a new adaptation evolved specifically within the research lab environment? While no one has studied the question in detail, we can find some clues by digging through the scientific archive.
Φ80 was first isolated in 1961 from a collection of clinical samples from Osaka University’s Department of Clinical Evaluation. The version of Φ80 studied by the Urbana-Champaign team is genetically identical to the original, Japanese variety, so any special adaptations must have occurred before 1961. While the phenomenon of P1 transduction was already known at the time, it was not yet used as a method for genome engineering, so it’s unlikely that the “hitchhiking” of molecular biology techniques was a new trait actively selected for.
All in all, it seems likely that Φ80’s ability to infiltrate biology labs is an exaptation: a trait that was originally selected for unrelated reasons, but was later repurposed in a different way after a change of environment. The genetic diversity of phages, after all, is so immense that it is unsurprising that one of them would turn out to be perfectly suited to life in the laboratory.
What Now?
Addressing the Φ80 crisis requires overcoming three obstacles: One is technological, one is social, and one is related to the very nature of biology as a science.
The technological challenge is the most tractable. For a long time, it was only practical to sequence short regions of the genome (using the Sanger method), so researchers would only PCR-amplify and sequence the particular locus they intended to modify. This can grant them great confidence that a strain carries the expected genotype, but it’s basically useless against a phage that inserts itself in a different region of the genome.
While whole-genome sequencing has become much more accessible, it doesn’t really solve the problem. First, it is more expensive than local Sanger sequencing. Second, short-read sequencing (e.g., Illumina) only generates short segments of DNA that must then be assembled on a computer. It is possible to use these short segments to stitch together the entire genome, much like a puzzle, but it’s slightly faster to align it to a reference genome, with the drawback that only sequences that map to the reference are considered. Sadly, the latter method will only detect a hidden phage if a researcher provides the phage genome as one of those references.
In other words, it would be feasible to find Φ80 through whole-genome sequencing, but it would require making two sub-optimal choices: using a slightly more expensive sequencing run and processing the results with a slightly slower algorithm. This is why, in practice, nobody does it. Fortunately, a third generation of sequencing can help: it’s becoming increasingly popular to sequence entire genomes using long-read (e.g., Nanopore) sequencing. This method reads DNA in long segments of thousands of nucleotides, making Φ80’s 46,000 nucleotides of foreign DNA much easier to spot. Hopefully, in the near future, this technique will help us identify Φ80 invasions more readily.
In the meantime, however, it remains difficult to identify Φ80 unless actively looking for it. This brings us to the social obstacles. For a microbiologist to check their strains, they need to be aware of the phage’s existence and dedicate time to checking for it.
Imagine being a fourth-year PhD student who finally managed to measure an interesting, statistically-significant, reproducible difference between two bacterial strains. Their hypothesis makes sense. They did everything right. As far as publication goes, their findings as they are would merit publication. The painful truth is that double-checking for Φ80 contamination is guaranteed to hurt their career. At best, they spend a few hours performing an additional experiment and find nothing; at worst, they discover that their results are artifactual and no longer publishable. There is no reward for discovering that your results are false positives.5
Finally, while Φ80 contamination may be unfortunate, there is something inevitable about this sort of setback given the inherent messiness and complexity of living organisms. So while Occam’s razor is a convenient heuristic, it does not actually work that well for biology; there is always a long tail of insanely complicated speculative possibilities that, every once in a while, turn out to materialize. A stealthy phage spreading from culture to culture? It may not be the most plausible explanation a priori, but it’s plausible enough. And if it’s not Φ80, it could still be any number of improbable but real bottlenecks: autoclave-resistant RNases, traces of phosphorus in the water, novel organelles, or parasites from an entirely unknown domain of life.
This is simply part of life as a biologist. In the 1990s, cognitive scientist Kevin Dunbar spent a year conducting ethnographic observations in a molecular biology lab. According to his notes, 40 to 60 percent of experimental results go against expectation, often for totally unknown reasons. As scientists push the frontier of knowledge, phenomena that were once considered mysterious become settled science, opening the way for new experimental techniques that themselves unearth new mysteries.
This actually makes me fairly hopeful about the future of biology. When people started experimenting with E. coli in the 1940s, they didn’t know a single thing about it. Since then, there has been considerable progress, and we now have at least some information about 3803 of its 4747 genes. The more we understand it, the less room there is left for unexpected artifacts creating misleading results and false interpretations, and the easier it becomes to make progress. Many outlandish oddities, higher-order emergent interactions, and bizarre edge cases loom ahead, but the fog is lifting.
In the end, we should embrace experimental artifacts and contamination not as setbacks to our research agenda, but rather as essential parts of the systems we study. Sometimes, as in the case of Φ80, the artifacts may even have a life of their own.
Antoine Vigouroux is a systems biologist. He writes about science and progress at Telescopic Turnip.
Thanks to Jay and Jasmine for feedback.
Cite: Vigouroux A. “How Φ80 Infiltrates Research Labs.” Asimov Press (2026). DOI: 10.62211/75eh-27rw
The technical term for a gene carried by a phage that is advantageous to the host bacteria is “moron.” In a remarkable display of temperance, I refrained from making a joke about it.
You might be wondering if there is any logical rule for naming phages. Short answer: not really. If you want to call your phage Belieber, no one can stop you. However, it is forbidden to name phages after Nicolas Cage.
On a technical note: P1 transduction does not use the “real” P1, but a mutant version called P1vir. The real P1 is lysogenic (like λ and Φ80), but for all practical purposes, the mutant behaves like a lytic phage (like T7). This changes nothing in the present story.
How do you get the antibiotic resistance gene there in the first place? That is a whole different story. In brief, pieces of DNA are usually assembled together in vitro using purified enzymes, then electroporated into the bacteria, where other enzymes incorporate it into the chromosomal genome. Incidentally, the most popular enzymes to do this are taken from phage λ. Much of what we can do in biotechnology has emerged downstream from figuring out how phages work and repurposing their tools.
I once discovered Φ80 myself in a strain I was working with. When I brought it up in a lab meeting, I don’t think any of the seasoned microbiologists in the room had ever heard of it. People joked that they’d rather not know about that, and as far as I can tell, nobody actually bothered to check their strains.


A classic method for detecting prophages is Southern blotting.
Love your writing style! Very aspirational.